PROBABILISTIC GRAPHICAL MODELS IN COMPUTATIONAL MOLECULAR BIOLOGY
1 MODELOS PROBABILISTICOS 11 MODELO BINOMIAL CIERTOS FENÓMENOS ALEATORIOSBEHAVIOR OF A PROBABILISTIC TABU SEARCH ALGORITHM FOR THE
CONTINUOUS VS DISCRETE PROCESSES THE PROBABILISTIC EVOLUTION OF SINGLE
PROBABILISTIC DAMAGE ANALYSIS OF OFFSHORE INSTALLATIONS SIROUS YASSERI KBR
PROBABILISTIC FORECAST KIYOHARU TAKANO CLIMATE PREDICTION DIVISION JMA 1
PROBABILISTIC GRAPHICAL MODELS IN COMPUTATIONAL MOLECULAR BIOLOGY
Probabilistic Graphical Models in Computational Molecular Biology
Probabilistic Graphical Models
in
Computational Molecular Biology
Pierre Baldi
University of California, Irvine
OUTLINE
INTRODUCTION: BIOLOGICAL DATA AND PROBLEMS
THE BAYESIAN STATISTICAL FRAMEWORK
PROBABILISTIC GRAPHICAL MODELS
APPLICATIONS
DATA COMPLEXITY AND COMPUTATIONAL PROBLEMS
Exponential data expansion.
Biological noise and variability. Evolution.
Physical and Genetic Maps.
Pairwise and Multiple Alignments.
Motif Detection/Discrimination/Classification.
Data Base Searches and “Mining”.
Phylogenetic Tree Reconstruction
Gene Finding and Gene Parsing.
Gene Regulatory Regions and Gene Regulation.
Protein Structure (Secondary, Tertiary, etc.).
Protein Function.
Genomics, Proteomics, etc.
MACHINE LEARNING
Machine Learning = Statistical Model Fitting.
Extract Information from the data automatically (inference) via a process of model fitting (learning from examples).
Model Selection: Neural Networks, Hidden Markov Models, Stochastic Grammars, Bayesian Networks.
Model Fitting: Gradient Methods, Monte Carlo Methods,…
Machine learning approaches are most useful in areas where there is a lot of data but little theory.
THREE KEY FACTORS
Data Mining/Machine Learning Expansion is fueled by:
Progress in sensors, data storage, and data management.
Computing power.
Theoretical framework: Bayesian Statistics, Probabilistic Graphical Modeling.
INTUITIVE APPROACH
Look at ALL available data, background information, and hypothesis.
Use probabilities to express PRIOR knowledge.
Use probabilities for inference, model selection, model comparison, etc. by computing POSTERIOR distributions and deriving UNIQUE answers.
DEDUCTION AND INFERENCE
DEDUCTION:
If AB and A is true,
then B is true.
INDUCTION:
If AB and B is true,
then A is more plausible.
BAYESIAN STATISTICS
Bayesian framework for induction: we start with hypothesis space and wish to express relative preferences in terms of background information (the Cox-Jaynes axioms).
Axiom 0: Transitivity of preferences.
Theorem 1: Preferences can be represented by a real number (A).
Axiom 1: There exists a function f such that
(non A)=f((A))
Axiom 2: There exists a function F such that
(A,B)=F((A), (B|A))
Theorem2: There is always a rescaling w such that P(A)=w((A)) is in [0,1], and satisfies the sum and product rules.
PROBABILITY AS DEGREE OF BELIEF
Sum Rule:
P(A|I) = 1-P(non-A|I)
Product Rule:
P(A,B|I) = P(A|I) P(B|A,I)
BayesTheorem:
P(A|B) = P(B|A) P(A) / P(B)
Induction Form:
P(Model|Data) = P(Data|Model) P(Model) / P(Data)
Equivalently:
P(Model|Data,I) = P(Data|Model,I) P(Model|I) / P(Data|I)
Recursive Form:
P(Model|D1,D2,…,Dn+1) = P(Dn+1|Model) P(Model|D1,…,Dn) / P(Dn+1|D1,…,Dn)
DIFFERENT LEVELS OF BAYESIAN INFERENCE
Level 1: Find the best model w*.
Level2: Integrate over models.
A non-probabilistic model is NOT a scientific model.
EXAMPLES OF NON-SCIENTIFIC MODELS
F=ma
E=mc2
etc…
These are only first-order approximations and do not “fit” the data (likelihood is zero).
Correction: (F+ F’) = (m+m’)(a+a’).
TO CHOOSE A SIMPLE MODEL BECAUSE DATA IS SCARCE IS LIKE SEARCHING FOR THE KEY UNDER THE LIGHT IN THE PARKING LOT.
MODEL CLASSES
BINOMIAL/MULTINOMIAL MODELS
NEURAL NETWORKS
MARKOV MODELS, KALMAN FILTERS
HIDDEN MARKOV MODELS
STOCHASTIC GRAMMARS
DECISION TREES
BAYESIAN NETWORKS
GRAPHICAL MODELS IS THE UNIFYING CONCEPT
LEARNING
MODEL FITTING AND MODEL COMPARISON
MAXIMUM LIKELIHOOD AND MAXIMUM A POSTERIORI
PRIORS
NON-INFORMATIVE PRIORS (UNIFORM, MAXIMUM ENTROPY, SYMMETRIES)
STANDARD PRIORS: GAUSSIAN, DIRICHLET, ETC.
LEARNING ALGORITHMS
Minimize -log P(M|D).
Gradient methods (gradient descent, conjugate gradient, back-propagation).
Monte Carlo methods (Metropolis, Gibbs sampling, simulated annealing).
Other methods: EM (Expectation-Maximization), GEM, etc.
OTHER ASPECTS
Model complexity.
VC dimension.
Minimum description length.
Validation and cross validation.
Early stopping.
Second order methods (Hessian, Fisher information matrix).
etc.
AXIOMATIC HIERARCHY
GAME THEORY
DECISION THEORY
BAYESIAN STATISTICS
GRAPHICAL MODELS
GRAPHICAL MODELS
Bayesian statistics and modeling leads to very high-dimensional distributions P(D,H,M) which are typically intractable.
Need for factorization into independent clusters of variables that reflect the local (Markovian) dependencies of the world and the data.
Hence the general theory of graphical models.
Undirected models reflect correlations: Random Markov Fields, Boltzmann machines, etc.
Undirected models are used for instance in image modeling problems.
Directed models reflect temporal and causality relationships: NNs, HMMs, Bayesian networks, etc.
Directed models are used for instance in expert systems.
Mixed Directed/Undirected Models and other variations are possible.
BASIC NOTATION
G=(V,E) = graph.
V = vertices, E = directed or undirected edges.
XI = random variable associated with vertex i.
XY = X and Y are independent.
XY|Z = X and Y are independent given Z
P(X,Y|Z)=P(X|Z) P(Y|Z)
N(i) = neighbors of vertex i.
Naturally extended to sets and to oriented edges.
“+” = children or descendants or consequences or future.
“–” = parents or ancestors or causes or past.
C+(i) = the future of i.
Oriented case: topological numbering of the vertices.
UNDIRECTED GRAPHICAL MODELS
Undirected models reflect correlations: Random Markov Fields, Boltzmann machines, etc.
Undirected models are used for instance in image modeling problems, statistical mechanics of spins, etc.
Markov properties are simpler. Global factorization is more complex.
MARKOV PROPERTIES
Pairwise Markov Property: Non-neighboring pairs Xi and Xj are independent conditional on all the other random variables.
Local Markov Property: Conditional on its neighbors, any variable Xi is independent of all other variables.
Global Markov Property: If I and J are two disjoint sets of vertices, separated by a set K, the variables in I and J are independent conditional on the variables in K.
Theorem: The 3 Markov properties above are equivalent. In addition, they are equivalent to the statement that the probability of a node given all the other nodes is equal to the probability of the node given its neighbors only.
GLOBAL FACTORIZATION
P(Xi | Xj : j in N(I)) are the local characteristics of the Markov random field. They uniquely determine the global distribution, but in a complex way.
The global distribution can be factorized as:
P(X1,…,Xn) = exp [-C fC(XC)] / Z.
fC = potential or clique function of clique C
maximal cliques: maximal fully interconnected subgraphs
DIRECTED GRAPHICAL MODELS
Directed models reflect temporal and causality relationships: NNs, HMMs, Markov Models, Bayesian Networks, etc.
Directed models are used, for instance, in expert systems.
Directed Graph must be a DAG (directed acyclic graph).
Markov properties are more complex. Global factorization is simpler.
MARKOV PROPERTIES
The future is independent of the past given the present
Pairwise Markov Property: Non-neighboring pairs Xi and Xj with i < j are independent, conditional on all the other variables in the past of j.
Local Markov Property: Conditional on its parents, a variable is independent of all the other nodes, except for its descendants (d-separation). Intuitively, i and j are d-connected if and only if either (1) there is a causal path between them or (2) there is evidence that renders the two nodes correlated with each other.
Global Markov Property. Same as for undirected graphs but with generalized notion of separation (K separates I and J in the moral graph of the smallest ancestral set containing I, J, and K.
GLOBAL FACTORIZATION
The local characteristics are the parameters of the model. They can be represented by look-up tables (costly) or other more compact parameterizations (Sigmoidal Belief Networks, NNs parameterization, etc.).
The global distribution is the product of the local characteristics:
P(X1,…,Xn) = i P(Xi|Xj : j parent of i)
BELIEF PROPAGATION OR INFERENCE
Basically a repeated application of Bayes rule.
TREES
POLYTREES (Pearl’s algorithm)
GENERAL DAGS (Junction Tree Algorithm, Lauritzen, etc.)
RELATIONSHIP TO OTHER MODELS
Neural Networks.
Markov Models.
Kalman Filters.
Hidden Markov Models and the Forward-Backward Algorithm.
Interpolated Markov Models.
HMM/NN hybrids.
Stochastic Grammars and the Inside-Outside Algorithm.
New Models: IOHMMs, Factorial HMMs, Bidirectional IOHMMs, etc.
APPLICATIONS
PROBABILISTIC RISK CRITERIA FOR NUCLEAR POWER PLANTS WEDNESDAY JANUARY
PROBABILISTIC SESMIC HAZARD ASSESSMENT FOR CHENNAI IGC 2009 GUNTUR
Tags: biology ===============================, graphical, biology, computational, molecular, models, probabilistic
- DR CHELLE L GENTEMANN [SHEHER] SENIOR SCIENTIST FARALLON INSTITUTE
- WETTABLE ACRES DETERMINATION CERTIFICATION NAME OF FACILITY FACILITY NUMBER
- VAN SEDNICE UPRAVNOG ODBORA DONETE SU SLEDEĆE ODLUKE
- [MIRTH1848] ERROR MESSAGE AFTER TRYING TO TRANSFER A FILE
- NR INTRARE FRB …………………… NR LICENTA FRB CERERE PENTRU
- 1 60 Jahre cjd in Elze das Christliche Jugenddorfwerk
- EL JUEGO DE LAS PATENTES II A TRAVÉS DE
- WOGA286 PÁGINA 2 OMPI S WOGA286 ORIGINAL INGLÉS FECHA
- 7 TEMA PALŪKANŲ NORMOS IR OBLIGACIJOS 20211101 2 PSL
- INFLUENCES OF CARBOHYDRATE PLUS AMINOACID SUPPLEMENTATION ON DIFFERING EXERCISE
- AL DHAYGHAMIAH AGRICULTURE CO (ADAC) WADI HALI AQUACULTURE PROJECT
- THANKSGIVING POINT WELCOME THE NASA DESTINATION STATION TO WATER
- 20175 HÁZIORVOSI KÉRDŐÍV TISZTELT HÁZIORVOS ASSZONYÚR! ALULÍROTT …………………………………………………………………………… NEVŰ
- 7TH GRADE LANGUAGE USAGE SENTENCES 3 NAME QUESTION
- 9 MUSIC FOR 20 NOTE ORGANS DANCES 406 JOSEF
- PUBLICADO EN EL REGISTRO OFICIAL NO 375 DE 12
- 16 LAMPIRAN 3(C) PERJANJIAN UTAMA SEWA RUANG BANGUNAN KANTIN
- VETERINARSKA STANICA VUKOVAR DOO V U K O V
- SZPITAL KLINICZNY IM KS ANNY MAZOWIECKIEJ UL KAROWA 2
- MOBİL UYGULAMA GİZLİLİK BİLDİRİMİ SÜRÜM 16 NISAN 2020
- DRAFT DOCUMENT – FOR CONSULTATION INCREASING PARTICIPATION IN FAITH
- TIPS FOR MAKING ART WITH YOUNG PEOPLE THE YOUNG
- 54MBPS 80211G WIRELESS PCI ADAPTER TRENDNETS TEW403PI 54MBPS WIRELESS
- CHURCH VIEW PRACTICE (YELLOW SUITE) PATIENT PARTICIPATION GROUP MEETING
- TITLE (TIMES NEW ROMAN 14 PT BOLD CENTRE ALIGNED)
- TC ONDOKUZ MAYIS ÜNİVERSİTESİ SAMSUN ( FORM17) MEZUNİYET TALEP
- A DUNA HÁZ TÁRSASHÁZ SZERVEZETI ÉS MŰKÖDÉSI SZABÁLYZATÁNAK 5
- DOKUMENTATION SCHÜLERTUTORIEN NAME TUTORIN NAME TUTANDIN FACH DATUM THEMA
- FORM A NOTIFICATION OF CHILD DEATH CDOP
- SUBJECT LINE 26 FEBRUARY 2014 NASA HELPING CALIFORNIA DEAL
 RENSTRA DINAS PERUMAHAN RAKYAT DAN KAWASAN PERMUKIMAN KABUPATEN SAMPANG
RENSTRA DINAS PERUMAHAN RAKYAT DAN KAWASAN PERMUKIMAN KABUPATEN SAMPANGCONFLICTOS DE INTERES DE LOS ARBITROS DEBER DE REVELACION
 CADASTRAMENTO DE SERVIDOR NO SISTEMA ONLINE S2ID 1 POR
CADASTRAMENTO DE SERVIDOR NO SISTEMA ONLINE S2ID 1 PORZNAK SPRAWY DOZ383162017 ZAŁĄCZNIK NR 6 DO SIWZ SZCZEGÓŁOWY
 AVTALE OM RETNINGSLINJER FOR OIDRETTENS
AVTALE OM RETNINGSLINJER FOR OIDRETTENS  HUMAN RESOURCES DIVISION DISCIPLINARY HEARING RECORD
HUMAN RESOURCES DIVISION DISCIPLINARY HEARING RECORD 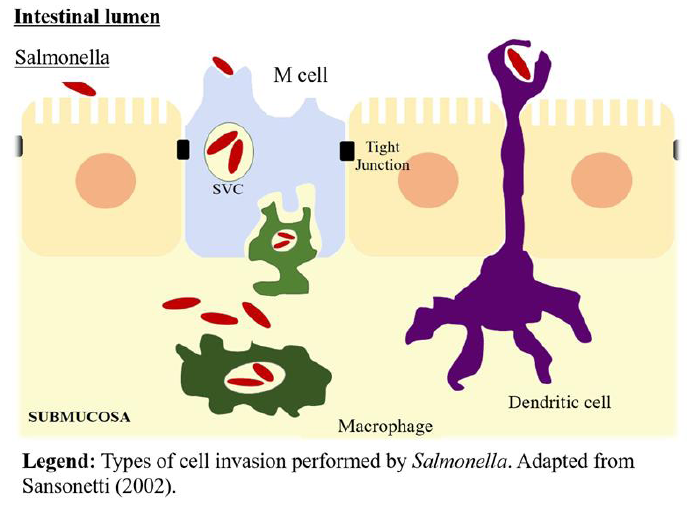 SALMONELLA SPP IN THE FISH PRODUCTION CHAIN AREVIEW SALMONELLA
SALMONELLA SPP IN THE FISH PRODUCTION CHAIN AREVIEW SALMONELLA R VIEUXFERRETTE ÉPUBLIQUE FRANÇAISE TÉL 0389404060 DÉPARTEMENT DU
R VIEUXFERRETTE ÉPUBLIQUE FRANÇAISE TÉL 0389404060 DÉPARTEMENT DU 16 E 17 LUGLIO MEZZANO DI PRIMIERO IN FESTA
16 E 17 LUGLIO MEZZANO DI PRIMIERO IN FESTA ATVIRO KONKURSO SĄLYGŲ 8 PRIEDAS ŽEMAITIJOS NACIONALINIO PARKO LAUKO
ATVIRO KONKURSO SĄLYGŲ 8 PRIEDAS ŽEMAITIJOS NACIONALINIO PARKO LAUKO DOSSIER DE PREMSA COMENÇA LA MARATÓ DE
DOSSIER DE PREMSA COMENÇA LA MARATÓ DEЗАГАШЕВ И ПРИЕМ «ЧТО? КТО? ГДЕ? КАК? КОГДА?» ЦЕЛЬ
 EL ESTUDIANTE EN EL CENTRO DE LA EDUCACIÓN A
EL ESTUDIANTE EN EL CENTRO DE LA EDUCACIÓN A EL METROSUR (1) PARA HACER ESTA FICHA TIENES QUE
EL METROSUR (1) PARA HACER ESTA FICHA TIENES QUEAYUDAS PARA QUE INSTITUCIONES ESPAÑOLAS INVITEN A CONFERENCIANTES FULBRIGHT
MODELLO DI SEGNALAZIONE DI CONDOTTE ILLECITE AI SENSI DELL’ART
 KSW 3307 KÁVOMLYNČEK VŠEOBECNÉ BEZPEČNOSTNÉ SMERNICE PRED POUŽITÍM TOHTO
KSW 3307 KÁVOMLYNČEK VŠEOBECNÉ BEZPEČNOSTNÉ SMERNICE PRED POUŽITÍM TOHTOFORM OF AUTHORIZING RESOLUTIONS FOR BORROWERS AS EVIDENCED BY
 3 GABINETE DE PRENSA Y COMUNICACIONES LEÓN 1 DE
3 GABINETE DE PRENSA Y COMUNICACIONES LEÓN 1 DE1 DE DICIEMBRE 2016 EMPRESA BONILLA SA DE CV